This page was produced as an assignment for Genetics 677, a undergraduate course at UW-Madison
CAPN3 Protein Homology
________________________________________________________________________________________________________
To find out how broadly CAPN3 is conserved, I searched NCBI Protein, Nucleotide, and HomoloGene for species with known capn3 homologs (keywords: capn3, canp3, calpain 3, calpain-3).
15 entries were found:
Bos taurus (cow)
Crisetulus griseus (chinese hamster)
Danio rerio (zebrafish)
Gallus gallus (chicken)
Heterocephalus glaber (naked mole rat)
Hippoglossus hippoglossus (atlantic halibut)
Macaca fascicularis (crab-eating macaque)
|
Macaca mulatta (rhesus macaque)
Mus musculus (mouse)
Oryctolagus cuniculus (rabbit)
Ovis aries (sheep)
Rattus norvegicus (rat)
Salmo salar (atlantic salmon)
Sus scrofa (wild boar)
Xenopus laevis (african clawed frog)
|
Discussion
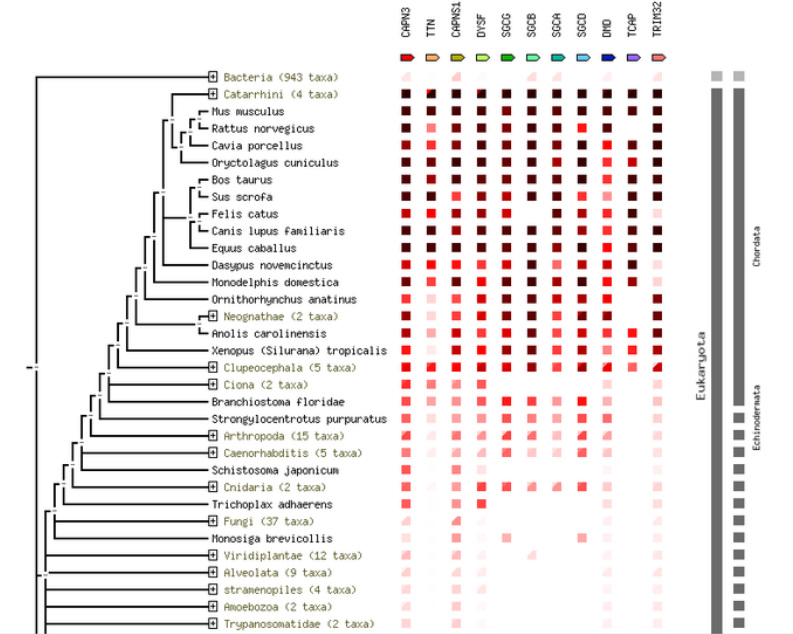
CAPN3, p94 is broadly conserved within species of the clade Euteleostomi and somewhat conserved beyond this clade, as also shown in the phylogenetic chart from STRING (Figure 1).
There were fewer entries for CAPN3 protein homologs than for homologous genes, and this may be due to incomplete characterization of the gene sequence. Many of the entries still hold PREDICTED or MODEL RefSeq status* and have not been experimentally verified.
*A brief explanation of how RefSeq status is determined can be found here
There were fewer entries for CAPN3 protein homologs than for homologous genes, and this may be due to incomplete characterization of the gene sequence. Many of the entries still hold PREDICTED or MODEL RefSeq status* and have not been experimentally verified.
*A brief explanation of how RefSeq status is determined can be found here
CAPN3 Protein Homology (from STRING)
________________________________________________________________________________________________________
|
References
[1] Dereeper A., Audic S., Claverie J.M., Blanc G. BLAST-EXPLORER helps you building datasets for phylogenetic analysis. BMC Evol Biol. 2010 Jan 12;10:8. [2] Dereeper A.*, Guignon V.*, Blanc G., Audic S., Buffet S., Chevenet F., Dufayard J.F., Guindon S., Lefort V., Lescot M., Claverie J.M., Gascuel O. Phylogeny.fr: robust phylogenetic analysis for the non-specialist. Nucleic Acids Res. 2008 Jul 1;36(Web Server issue):W465-9. Epub 2008 Apr 19. [3] Edgar RC. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, Mar 19;32(5):1792-7. [4] Jensen LJ, et al. STRING 8--a global view on proteins and their functional interactions in 630 organisms. Nucleic Acids Res. 2009 Jan;37(Database issue): D412-6. Epub 2008 Oct 21. |
_[5] Castresana J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol. 2000, Apr;17(4):540-52.
[6] Guindon S., Gascuel O. A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood. Syst Biol. 2003, Oct;52(5):696-704. [7] Anisimova M., Gascuel O. Approximate likelihood ratio test for branchs: A fast, accurate and powerful alternative. Syst Biol. 2006, Aug;55(4):539-52. [8] Di Tommaso P, Moretti S, Xenarios I, Orobitg M, Montanyola A, Chang JM, Taly JF, Notredame C. T-Coffee: a web server for the multiple sequence alignment of protein and RNA sequences using structural information and homology extension. Nucleic Acids Res. 2011 Jul;39(Web Server issue):W13-7. Epub 2011 May 9. |